RESEARCH ARTICLE
Prevalence of Methicillin Resistant and Virulence Determinants in Clinical Isolates of Staphylococcus aureus
Manjunath Chavadi1, Rahul Narasanna1, Ashajyothi Chavan1, Ajay Kumar Oli2, Chandrakanth Kelmani. R1, *
Article Information
Identifiers and Pagination:
Year: 2018Volume: 10
First Page: 108
Last Page: 115
Publisher ID: TOIDJ-10-108
DOI: 10.2174/1874279301810010108
Article History:
Received Date: 5/5/2018Revision Received Date: 18/7/2018
Acceptance Date: 28/7/2018
Electronic publication date: 13/08/2018
Collection year: 2018

open-access license: This is an open access article distributed under the terms of the Creative Commons Attribution 4.0 International Public License (CC-BY 4.0), a copy of which is available at: https://creativecommons.org/licenses/by/4.0/legalcode. This license permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Introduction:
Methicillin-resistant Staphylococcus aureus (MRSA) is the major threat that is a result of the uncontrolled use of antibiotics causing a huge loss in health, so understanding their prevalence is necessary as a public health measure.
Objective:
The aim of this study was to determine the prevalence of methicillin-resistant MRSA and virulence determinant among associated S. aureus from the clinical samples obtained from various hospital and health care centers of the Gulbarga region in India.
Materials and Methods:
All the collected samples were subjected for the screening of S. aureus and were further characterized by conventional and molecular methods including their antibiotic profiling. Further, the response of methicillin antibiotic on cell morphology was studied using scanning electron microscopy.
Results:
A total 126 S. aureus was isolated from the clinical samples which showed, 100% resistant to penicillin, 55.5% to oxacillin, 75.3% to ampicillin, 70.6% to streptomycin, 66.6% to gentamicin, 8.7% to vancomycin and 6.3% to teicoplanin. The selected MRSA strains were found to possess mecA (gene coding for penicillin-binding protein 2A) and femA (factor essential for methicillin resistance) genetic determinants in their genome with virulence determinants such as Coagulase (coa) and the X region of the protein A (spa) gene. Further, the methicillin response in resistant S. aureus showed to be enlarged and malformed on cell morphology.
Conclusion:
The molecular typing of clinical isolates of S. aureus in this study was highly virulent and also resistant to methicillin; this will assist health professionals to control, exploration of alternative medicines and new approaches to combat Staphylococcal infections more efficiently by using targeted therapy.
1. INTRODUCTION
The fight between humans and tiny creatures called microorganisms (especially the pathogenic microorganisms) is from ancient times. The discovery of first antibiotic penicillin by Alexander Fleming (1928) started the golden era of
antibiotic, where everything seemed to be controlled well, but soon because of widespread and uncontrolled use of penicillin, the golden age came to an end as microorganisms became resistant to them. Not only penicillin but various other antibiotics have also become resistant, which have led to high morbidity and mortality [1]. Among various drug-resistant microorganisms, Methicillin-resistant forms of Staphylococcus aureus (MRSA) are a major threat causing a wide range of infections from the skin to pneumonia [2, 3]. S. aureus resistant to penicillin resulted in the introduction of methicillin, however, the preface of methicillin marks the onset of the second wave of resistance. The first reports on an S. aureus strain that was resistant to methicillin were published in 1961 [4], till now its prevalence has increased drastically, causing various health problems worldwide [5, 6]. Resistance to methicillin mediated via the mec operon, part of the Staphylococcus Cassette Chromosome mec (SCCmec) and conferred by the mecA gene, which codes for an altered penicillin-binding protein (PBP2a or PBP2') that has a lower affinity for binding β-lactams (penicillins, cephalosporins, and carbapenems). This allows for resistance to all β-lactam antibiotics and obviates their clinical use during MRSA infections, such as glycopeptide vancomycin is often deployed against MRSA.
S. aureus expresses many potential virulence factors such as a capsular polysaccharide, protein A and Leukocidin that have the potential to interfere with the host defense mechanisms. Capsular Polysaccharide impedes phagocytosis, Protein A binds to immunoglobulin G molecules by the Fc region, causing damage to the immune mechanism. S. aureus expresses several different types of protein toxins which damage the membranes of erythrocytes and leukocytes. The toxins are of various types, such as membrane-damaging toxins (α-toxin, β-toxin, δ-toxin and γ-toxin and leukocidin), superantigens, epidermolytic and other extracellular proteins such as coagulase and staphylokinase [7].
Treatment of Staphylococcal infections with antibiotics is becoming increasingly challenging for clinicians because of the widespread increase in multidrug resistance in S. aureus isolates [8]. However, at the same time, the pace of development of new antibiotics has beenslow, resulting in the shortage of novel classes of antibacterial compounds with which to eliminate multidrug-resistant pathogens [9, 10]. This leads to portray that studies for understanding its prevalence are required if public health concern is to be improved. In this regard, the present study is an attempt towards understandings (a) the prevalence of MRSA in the Gulbarga region, India. (b) The molecular typing of clinical isolates of S. aureus to understand its virulence, antibiotic resistance profiles and the response of methicillin to various forms of S. aureus.
2. MATERIALS AND METHODS
2.1. Isolation and Identification of S. aureus
In this study, a total 126 strains of S. aureus were isolated from a different clinical sample like pus, blood, and other exudates collected from different hospitals and healthcare centers of Gulbarga region India, during the period from March 2016 to April 2017. The preliminary identification of S. aureus was done using mannitol salt agar which was detected by changes in the color of the medium from red to yellow due to mannitol fermentation. Further, the S. aureus was identified based on cultural, morphological, and biochemical tests. Standard strain S. aureus (MTCC 96) was obtained from the Microbial Type Culture Collection and Gene Bank center, Chandigarh, India and used as a control.
2.2. Antibiotic Susceptibility Test
The phenotypic detection of MRSA was done by an antibiotic susceptibility test performed using antibiotics discs which include penicillin, ampicillin, gentamicin, streptomycin, kanamycin, and methicillin (Hi-media, Mumbai) used as recommended by the Clinical and Laboratory Standards Institute (CLSI) guidelines [11] by the agar disc diffusion method.
2.3. Minimum Inhibitory Concentration (MIC)
The MIC is defined as the lowest antibiotic concentration with no visible growth. Based on the MIC values i.e. S. aureus with an MIC in the range of 0.5 - 4.0 µg/mL and MIC above 16 µg/mL with respect to oxacillin antibiotic was considered as methicillin sensitive S. aureus (MSSA) and MRSA, respectively. The MIC was determined by agar dilution method using the CLSI guidelines on plates of Muller-Hinton agar containing serial gradient dilution with oxacillin (4 -1024 µg/mL Sigma-Aldrich). Bacterial suspension equivalent to 0.5 MacFarland standard, was inoculated on the surface of Mueller Hinton plates, with the help of sterile cotton swabs and incubated overnight at 37°C for 24 h for visible growth. For determination of vancomycin resistance, the gradient plates of Mueller-Hinton agar were prepared with vancomycin (2-128 µg/mL Sigma-Aldrich). 0.5 McFarland equivalent inoculum prepared using 18-24 h old culture was put on gradient plates. The plates were incubated overnight at 35°C for 24 h before assessing the visible growth.
2.4. Response of Antibiotic on S. aureus Morphology
The S. aureus morphology was determined using scanning electron microscopy (S-200C scanning electron microscope). The isolated MRSA, MSSA and standard control MTCC 96 were grown on Brain Heart Infusion broth (BHI). The bacterial cells from each culture were recovered by centrifugation at 6000 RPM for 5 minutes and the cells were washed twice with potassium phosphate buffer (50 mM, pH 7.0). Bacterial cells were fixed by immersing in 2.5% glutaraldehyde in potassium phosphate buffer (50 mM, pH 7) for overnight at 4°C. Then the specimens were washed twice with buffer and dehydrated by ethanol series (v/v) ranging from 30%, 40%, 50%, 60%, 70%, 80%, 90% to 100%. For SEM analysis, all the specimens were dried to the critical point and coated with gold. The cell volume and surface area obtained from the SEM photograph were directly measured and calculated by using the following equations;
V (µ m3) = 4/3 πr3,
a (µ m2) = 4πr2;
Where “r” is the radius, “V” is the volume and “a” is the surface area. The average cellular volume and surface area were calculated by using 30 individual bacteria per population. Cells showing deformations were not considered. The mean values were calculated from SEM photographs by taking 30 bacteria per population. Statistics were calculated using the ANOVA [12, 13].
2.5. Preparation of Chromosomal DNA
Cells from an overnight culture in BHI broth collected by centrifugation were suspended in lysis buffer (phosphate-buffered saline [PBS] containing 1% Sodium Dodecyl Sulfate [SDS] and 100 µg/mL proteinase K). The cell suspension was incubated at 37°C for 1 h, and an equal volume of phenol: chloroform (1:1) mixture was added to the cell suspension and vortexed. The samples were centrifuged, and the aqueous phase was transferred to a fresh tube. The DNA was precipitated by the addition of 100 µL of 3M sodium acetate and 3 volume of chilled absolute ethanol; the DNA pellet was washed twice with chilled 70% alcohol, air-dried, and suspended in 50 µL of TE buffer (10 mM Tris-HCl [pH 8.0], 1 mM EDTA) [14].
2.6. PCR for Detection of coa, spa, femA and mecA
For all molecular studies, five highly resistant isolates designated as VMRSA102 VMRSA26, MRSA09 MRSA10, and MRSA20 were selected. The identification of Coagulase (coa) and the X region of protein A (spa) gene was amplified by PCR using the forward primer 5′-CAAGCACCAAAAGAGGAA-3′ and reverse primer 5′-CACCAGGTTTAACGACAT -3′ for coa gene and forward primer 5′- AGACGATCCTTCGCTCAGC -3′and reverse primer 5′- GCTTTTGCAATGTCATTTACTG -3′ for the spa. The PCR was performed according to the previously described procedure [14, 15]. The presence of mecA (gene coding for penicillin binding protein 2A) and femA (factor essential for methicillin resistance) genes were amplified by PCR using the forward primer 5′-ACTGCTATCCACCCTCAAAC-3′ and reverse primer 5′-CTGGTGAAGTTGTAATCTGG-3′ for mecA and forward primer 5′-AAAAAAGCACATAACAAGCG-3′and reverse primer 5′-GATAAAGAAGAAACCAGCAG-3′ for femA. The PCR was performed according to the previously described procedure [14]. The amplified products were purified and run on 1% agarose gel containing ethidium bromide and the bands were observed under UV transilluminator to record the number of bands and their positions.
3. RESULTS
3.1. Antibiotic Profile and Resistance Rate Among the S. aureus Isolates
In this study, a total of 126 S. aureus were isolated from the clinical samples collected from various hospitals and healthcare centers of Gulbarga region, India. All S. aureus isolates were subjected to antibiotic susceptibility testing and results are shown in Tables 1 & 2. Among 126 S. aureus strains, 53.9% of isolates were found to be methicillin-resistant, which were mainly isolated from pus and blood samples. Antibiotic-resistant profile of S. aureus isolates showed 100% resistance to penicillin, 55.5% to oxacillin, 75.3% to ampicillin, 70.6% to streptomycin and 66.6% to gentamicin.
| Clinical Sample | No. Of Strains Isolated | No. of MRSA (%) |
|---|---|---|
| Pus | 56 | 38 |
| Urine | 24 | 09 |
| Blood | 30 | 16 |
| Sputum | 11 | 03 |
| Miscellaneous | 05 | 02 |
| Total | 126 | 68 |
| Antibiotics | No. Of Resistance Strains | Percentage (%) |
|---|---|---|
| β-lactam | ||
| Penicillin(10units) | 126 | 100 |
| Oxacillin(1mcg) | 70 | 55.5 |
| Methicillin (5mcg) | 68 | 53.9 |
| Aminoglycosides | ||
| Gentamicin (10 mcg) | 84 | 66.6 |
| Streptomycin (10 mcg) | 89 | 70.6 |
| Ampicillin (10 mcg) | 95 | 75.3 |
| Glycopeptides | ||
| Vancomycin | 11 | 8.7 |
3.2. Minimum Inhibition Concentration(MIC)
Out of 126 strains of S. aureus, 68 were MRSA and 58 were MSSA. Further based on the MIC results, MRSA isolates were classified as low-level oxacillin resistant (n = 30) strains (MICs ranged from 16 to >128 µg/mL) and high-level oxacillin-resistant strains (n =38; MIC >128 µg/mL). The MIC for 11 of 128 isolates (8.7%) for vancomycin was in the range of 16-64 mg/l indicating that those were vancomycin-resistant (VRSA) and the remaining 117 isolates showed an MIC range ≤ 2 mg/l indicating that all were sensitive to vancomycin.
The highest and lowest MICs for MRSA isolates were recorded as 720µg/mL for the isolates VMRSA102 and 16µg/mL for MRSA10 isolates. High-level oxacillin resistant isolates VMRSA102, VMRSA26, MRSA09 and Low-level oxacillin-resistant strains MRSA10 and MRSA20 (Table 3) have been further selected for typing of coa, spa, femA, and mecA.
| Isolate Number | MIC of Oxacillin (µg/ml) | coa- PCR typea | spa-PCR Typeb | mecA PCRc | femA PCRd |
|---|---|---|---|---|---|
| VMRSA102 | 720 | 2 | 1 | + | + |
| VMRSA26 | 512 | 2 | 1 | + | + |
| MRSA09 | 256 | 2 | 1 | + | + |
| MRSA20 | 128 | 1 | 1 | + | + |
| MRSA10 | 16 | 1 | 1 | + | + |
b Numbers of the patterns correspond to those of PCR amplification products of the spa gene.
c,d Typing of mecA and femA, results listed as positive (+).
3.2.1. The Response of Antibiotic on S. aureus Morphology
In order to evaluate the morphology of S. aureus, SEM was performed and summarized (Table 4). The results showed that MRSA cells have altered their morphology with respect to MSSA and standard S. aureus MTCC 96. The cell morphology of both MSSA and MTCC 96 cells was apparently normal (Fig. 1a, b), but enlarged and malformed in the MRSA cells (Fig. 1c).
| Bacterial Strains |
Radiusa (µm) | Volumeb(µm3) | Surface Area (µm2) |
|---|---|---|---|
| MTCC 96 | 0.346±0.0158 | 0.177±0.0239 | 1.49±0.151 |
| MSSA | 0.343±0.023 | 0.164±0.025 | 1.48±0.265 |
| MRSA | 0.453±0.0316 | 0.399±0.076 | 2.59±0.354 |
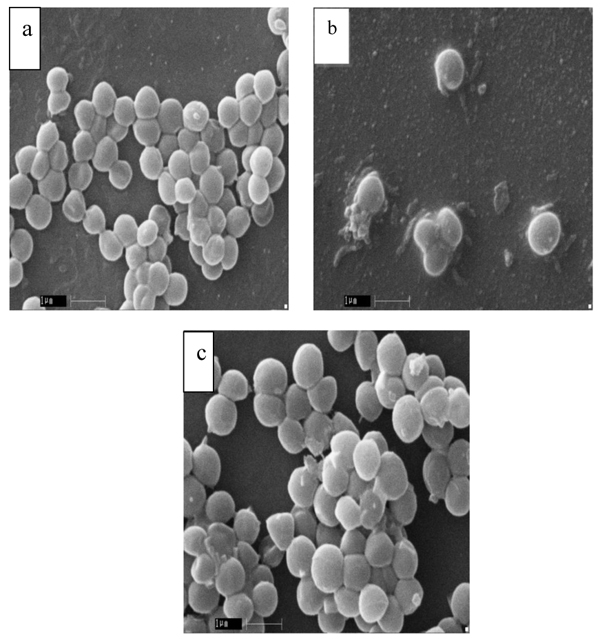 |
Fig. (1). Scanning electron micrographs of S. aureus cells (a) MTTC 96 (b) MSSA (c) MRSA. Bar 1 µm. |
3.3. Detection of coa, spa, femA and mecA
The production of coagulase is an important feature used worldwide for the identification of virulence factors of S. aureus. The five selected isolates VMRSA102, VMRSA26, MRSA09 MRSA10, and MRSA20 were coagulase positive. Although the five isolates were coagulase positive, the coagulase gene can be differentiated from each other on the basis of the presence of PCR products and their size. A single PCR product from MRSA10 and MRSA20 strains was amplified by the amplicon size of 270bp, while two PCR products were amplified from VMRSA102, VMRSA26 and MRSA09 strains and their sizes approximately ranged from 270 and 500 bp (Table 3).
The polymorphic X-region was used for the detection of protein A gene (spa). The five strains VMRSA102, VMRSA26, MRSA09, MRSA10, and MRSA20 were tested positive for amplification of protein A and found to have a single amplification PCR product of 300bp (Table 3). The five selected isolates VMRSA102, VMRSA26, MRSA09, MRSA10, and MRSA20 were found to be mecA and femA positive. The size of the amplified products for mecA and femA was 130 bp and 170 bp, respectively (Table 3).
4. DISCUSSION
In fact, with the national rise in the incidence of MRSA infection in clinical settings and hospitals in India, the incidence of MRSA drastically increased from 20 to 32.8% in 2000 to 79.6% in 2011 [16-18]. This high incidence of S. aureus infection has been significant to correlate with enhanced epidemiology and pathogenicity of Methicillin-Resistant S. aureus (MRSA). However, the prevalence rate of MRSA is 35% with low isolation rate [8], but from the same locality, our study shows high incidence rate of 56.81% MRSA in samples isolated from hospitals and diagnostic clinical samples of Kalaburagi region, Karnataka, India. The increased rate of the prevalence may be due to the inadequate use of antibiotics, lack of awareness and unethical treatment before coming to the hospital which have all been contributing factors. On another hand, there may be a chance of new strains emerging with MDR in nature through horizontal gene transfer.
The changes in S. aureus cell morphology are possibly owing to the adaptive response to overcome the adverse environmental conditions mainly occurring due to antibiotic stress in this study and this response has already been reported with several bacterial species [19]. The change in the morphology may be due to antibiotic stress; with an increase in the cell size, the relative contact surface reduces and consequently the attachable surface for organic (antibiotic) compounds. Therefore, bigger cells can tolerate the stress conditions better than normal cells of the same species [20, 13].
The coagulase production is mainly used for the identification of S. aureus. The PCR amplification of coagulase revealed a single amplified product in the majority of clinical isolates of S. aureus. The strain VMRSA102, VMRSA26, and MRSA09 gave two amplified PCR products, strongly suggesting that these S. aureus isolates have more than one allelic form of the coagulase gene and have a high level of MIC for oxacillin antibiotic, as compared to a single PCR product obtained from MRSA10 and MRSA 20 which has a low level of MIC for oxacillin antibiotic. The previous studies have also shown more than one to eight different amplification products for coagulase gene, this is mainly due to varying numbers of tandem repeats in the coa gene. Thus, our results moderately correlate with the earlier reports [21].
The protein A is one of the major criterions used for the identification of S. aureus. The X region of the spa gene consists of a variable number of tandem repeats sequences which differ among S. aureus isolates both in their number and in the location [21]. The five selected isolates VMRSA102, VMRSA26, MRSA09, MRSA10 and MRSA20 gave a single amplified PCR product, this is mainly because of the lack of allelic form of the protein A gene and the result correlates with an earlier report [22].
The MRSA isolated in the present study was resistant to ≥ 05 antibiotics belonging to a different class of antibiotics including β-lactamase, aminoglycosides, and glycopeptides. The reason for multidrug resistance is the mechanism for the creation of highly resistant subclones which would be the transposition of mobile genetic elements such as integrons, transposons and even larger resistant gene clusters and chromosomal cassettes. These elements may either have evolved separately carrying the same genes such as SCCmec elements in MRSA [14]. Evidence suggests that problem may likely be due to prolonged and repeated usage of methicillin against S. aureus infection, which might result in multidrug resistance to other antibiotics [23].
The five selected isolates VMRSA102, VMRSA26, MRSA09, MRSA10, and MRSA20 were confirmed to acquire the presence of mecA and femA gene. The mecA gene is associated with low affinity of penicillin-binding protein, PBP2a and is highly conserved in Staphylococcus species, so it is possible to detect methicillin-resistant S. aureus by detection of these genes. The reason for selecting mecA gene is considered as gold standard and hallmark for the identification of MRSA strains [19]and it is associated with drug resistance determinant [14], while femA gene is associated with the high-level expression of methicillin resistance and involved in the metabolism of cell wall synthesis. Furthermore, femA cooperates with mecA for the expression of β-lactam resistance, and appears to be a unique feature of S. aureus, which serves as an identification marker of species [24].
CONCLUSION
The rationale for targeting only methicillin resistance among S. aureus isolates is to considerably increase prevalence in the clinical sample in recent days, therefore, our study would beneficially help assist and guide beta-lactam therapy, especially in the case of MRSA and multiresistant staphylococcal variants using molecular methods to characterize virulence determinants.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
This research was supported/partially supported by Indian Council of Medical research, No. 80/860/2014-ECD-I. We thank our colleagues Dr. Rajeshwari and Dr. Ajay Kumar Oli who provided insight and expertise that greatly assisted the research, during the progress of this work.








