REVIEW ARTICLE
A Global Overview of β-lactam Resistance Genes in Klebsiella pneumoniae
Yone da Silva1, Rafaela Ferrari3, Victor Augustus Marin2, Carlos Adam Conte Junior1, 3, 4, *
Article Information
Identifiers and Pagination:
Year: 2019Volume: 11
First Page: 22
Last Page: 34
Publisher ID: TOIDJ-11-22
DOI: 10.2174/1874279301911010022
Article History:
Received Date: 13/03/2019Revision Received Date: 17/05/2019
Acceptance Date: 28/05/2019
Electronic publication date: 30/06/2019
Collection year: 2019

open-access license: This is an open access article distributed under the terms of the Creative Commons Attribution 4.0 International Public License (CC-BY 4.0), a copy of which is available at: https://creativecommons.org/licenses/by/4.0/legalcode. This license permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Klebsiella pneumoniae is a gram-negative bacillus of the Enterobacteriaceae family, commonly associated with nosocomial infections. This pathogen is a serious public health problem as some of its strains are resistant to about 95% antimicrobials of the pharmaceutical market. This resistance is promoted by the production of the β-lactamase extended spectrum (ESBL) enzymes, one of the major causes of therapeutic failure. This review evaluated the incidence and distribution of resistance genes from Klebsiella pneumoniae to β-lactams worldwide. Our study was conducted with the subject the organism K. pneumoniae and β-lactamic resistance. The most reported genes were blaSHV-12, blaCTX-M-2 and blaSHV-5; with blaSHV-12 being the most described. The last two were present in all continents, characterizing its cosmopolitan profiles. The greatest genetic diversity was observed in the Asian and Oceania, where 41 different genes were isolated. Additionally, our review points out the coexistence of different classes of β-lactamases in a single bacterial isolate. Finally, knowledge of mechanisms associated with resistance of K. pneumoniae is of great public interest and the verification of resistance genes shows a variation over time and location highlights the importance of evaluating the mechanisms or strategies by which these variations occur.
1. INTRODUCTION
Klebsiella pneumoniae is a microorganism found in humans and other mammals, colonizing intestine, skin, nasopharynx and, several environmental niches. A constituent of the human intestinal microbiota, this bacterium, considered harmless, has gained notoriety by being identified as an opportunistic pathogen. This is responsible for a variety of diseases in humans and animals, such as urinary tract infection, pneumoniae, intra-abdominal infection, bloodstream infection, meningitis, pyogenic liver abscess and is a prominent nosocomial pathogen [1, 2]. Moreover, the occurrence and global spread of hypervirulent and MDR (multidrug-resistant) K. pneumoniae are increasingly reported in community-acquired and nosocomial infections [3], and the use of antimicrobials in the treatment of infections has become progressively more difficult [4, 5]. However, despite there has been an increase in the number of vulnerable populations to MDR pathogens, in addition to a renewed interest in the Klebsiella pneumoniae, many questions about this pathogen are still unknown, as to how resistance genes spread out and how they reach humans [6].
Over time, bacteria have developed several mechanisms by which they can withstand antimicrobials, rendering the action of these substances ineffective [3]. The resistance mechanisms of a gram-negative bacterium include: efflux pump; membrane permeability; production of β-lactamases; modification of antibiotic targets; and, acquisition of alternative metabolic pathways to those inhibited by the antibiotic [4, 6]. One of the most important mechanisms of resistance in bacteria is the Extended Spectrum β-lactamase (ESBL) production [7]. The first β-lactamase with the ability to hydrolyze β-lactam was described in 1983 in a K. ozaenae isolated in Germany and designated as SHV-2 (Sulfhydryl variable) [8, 9]. However, the initial outbreaks were related only in 1985 in France and the late ‘80s in the United States. Currently, over one hundred and fifty different ESBLs types have been identified. These classes of enzymes have a distinct spectrum of action and are classified as: Extended Spectrum β-lactamase (ESBL); Metalo-β-lactamase (MBL); and, β-lactamase Class C (AmpC) [8].
The definition of ESBLs is not consensual, however, some authors propose that an ESBL is any β-lactamase, generally acquired and not inherent in a species, that hydrolyzes rapidly or confers resistance to oxyimino-β-lactam chain, broad-spectrum cephalosporins, penicillins, and monobactams and is inhibited by clavulanate, sulbactam and tazobactam [10, 11]. Since the discovery of these enzymes, an increase of K. pneumoniae resistant to several antimicrobials has been reported such as aminoglycosides and even carbapenems [12, 13]. These latter are considered to be one of the most important antibiotics classes in the fight against ESBL-producing microorganisms due to their stability against the hydrolytic activity of this enzyme and also for the ability to penetrate the bacterium [14].
β-lactam antibiotics interact with diverse groups of bacterial enzymes. The enzymes, such as TEM-1 (Temoniera), TEM-2 and SHV-1, encoded by plasmids and widely distributed among enterobacteria, can hydrolyze a wide variety of penicillins and third generation of cephalosporins, which were originally developed as drugs capable of overcoming bacterial resistance. These enzymes have emerged as one of the most successful resistance mechanisms, which limit therapeutic options for treating various human infections, including Klebsiella [15, 16].
Given the therapeutic limitations caused by resistance mechanisms, several research groups have followed the distribution and susceptibility profile of the genus Klebsiella spp., to monitor possible outbreaks and set up the sensitivity profile in different regions [12, 13]. Based on the global dissemination of ESBL-producing Klebsiella pneumoniae and limitations of the current treatments, the implementation of prevention and control measures has become extremely important [13].
Considering the recognition of the genus Klebsiella spp. as an important pathogen involved in infections, its variations in the antibiotic resistance profile, and its emerging mechanisms of resistance, reinforces the need to know the local or regional characteristics of this genus, as well as the evolution of its resistance. Thus, monitoring of the resistance and prevalence of antimicrobial resistance determinants are urgently needed and may contribute to better targeting and use of antimicrobials. In this context, the present review focuses on the main aspects related to the distribution of resistance genes to β-lactam antibiotics associated with Klebsiella pneumoniae found in the scientific literature.
2. MATERIALS AND METHOD
The searches were conducted between February and June of 2018 in online health databases, and date delimitation was set as between 2000 and 2018. Editorials, letters, and Ph.D. theses were excluded. The following databases were used for the research: PubMed, SCOPUS, Science Direct, Latin American and Caribbean Literature in Health Sciences (LILACS), Virtual Electronic Scientific Library (SCIELO) and Portal of Periodicals of the Coordination for the Upper Level (Capes Newspapers). Additionally, the descriptors to perform the searches were: Klebsiella pneumoniae, Klebsiella pneumoniae resistance factors, Klebsiella pneumoniae with resistance to β-lactam antimicrobials, broad-spectrum β-lactamases, and extended-spectrum β-lactamase. There was no restriction of sample size for evaluation of publications and the articles selected were only in the English language. Exclusion criteria were: studies that exclusively investigated resistance to non-β-lactam antibiotics and non-human studies. Some studies considered essential to compose the review not included in any of the research bases were added (i.e., articles addressing pathologies in humans, characteristics of the bacterium and, some studies published earlier than the year 2000 due to the importance of it to a specific country and to better contextualize a finding).
3. RESULTS AND DISCUSSION
The descriptors and databases used allowed the identification of 84 scientific articles that fully met the inclusion criteria of the study. It should be noted that several studies found despite the treatment of β-lactam antibiotic resistance associated with K. pneumoniae did not present all relevant information for our review, e.g., resistance genes and/or sites where the microorganism was isolated, which consequently had to be excluded. Finally, not all the countries that constitute the different continents were considered since until the conclusion of our review no results were published between 2000 and 2018.
3.1. Antimicrobial Resistance
Antimicrobial resistance is a growing worldwide iatrogenic complication, which currently constitutes a major public health problem and is responsible for a significant increase in patients' morbidity and mortality [10, 17].
As commented above, the Enterobacteriaceae family has several mechanisms of resistance recognized. This group has also received special attention in recent years as being among the most isolated microorganisms in the laboratory routine [12]. Belonging to this family, one of the important pathogens for humans, with well-known and powerful mechanisms of resistance, is the genus Klebsiella spp [12, 13].
In recent years, increased antibiotic resistance in Enterobacteriaceae has become a major concern worldwide. Although β-lactam antibiotics have been widely used as the first treatment of severe infections, resistance to these drugs due to the acquisition of resistance genes has emerged and spread worldwide since the early 2000s, thus increasing concern about the occurrence of infections of hospital origin where these bacteria are widespread [18].
The progression of resistance of Klebsiella pneumoniae to antibiotics has caused great concern since the 1980s with the emergence of K. pneumoniae ESBL-producing [19]. Since them, other β-lactamases were discovered, for example, the carbapenemase. This enzyme was discovered in the United States in 2001 in K. pneumoniae and has expanded remarkably, being able to confer resistance to all β-lactams, even those associated with β-lactamase inhibitors [20]. Another important β-lactamase is the MBL, which degrades all β-lactams except aztreonam (monobactam) in vitro, whereas AmpC, in turn, can hydrolyze penicillins, monobactams, and cephalosporins until the third generation [21].
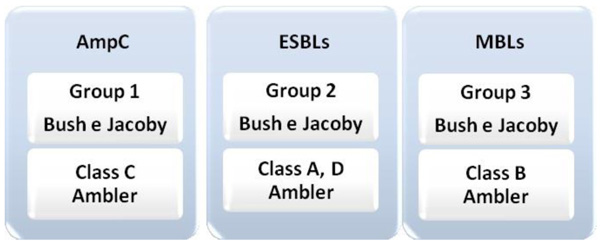
The β-lactamase type enzymes can be classified into two distinct ways, the molecular classification of Ambler [22] and the functional classification Bush & Jacoby [23]. The first classifies the β-lactamases into four classes (A, B, C, and D), grouped according to the similarity between the amino acid sequences. Classes A, C, and D are considered serine β-lactamase because they have the site of action composed of serine. Class B, also called MBL, uses zinc (Zn) as a cofactor [24].
The most current classification is from Bush & Jacoby, based on the activity of the enzyme, that is, on functional groups and correlates the properties of each enzyme with the resistance profile of each isolate [23, 24]. In this way, enzymes are classified into three groups: group 1, which is the group of cephalosporinases. Enzymes belonging to molecular class C, from Ambler, which are more active against cephalosporins and are not inhibited by β-lactamase inhibitors, such as clavulanic acid and tazobactam. Group 2 comprises the serine β-lactamases, enzymes belonging to group A and D, of Ambler, represented by the largest group of β-lactamases. It is divided into subgroups (2a and 2b) according to the action spectrum of each enzyme. Group 3 includes the enzymes that require the zinc ion at its site of action to produce antimicrobial activity (Fig. 1). They are the MBLs that can hydrolyze carbapenems and are inhibited by ion chelators, such as EDTA [23, 24].
The AmpC-type β-lactamases are chromosomal β-lactamases but can be found on both the bacterial chromosome and inserted into mobile plasmids. There are various types of plasmid-mediated AmpC β-lactamases: the CMY, MIR, MOX, LAT, FOX, DHA, ACT, ACC and CFE. They are capable of hydrolyzing penicillins, monobactams, and cephalosporins of up to the third generation, with resistance to cefoxitin being the main marker of AmpC expression. AmpC-producing bacteria are commonly also resistant to non-β-lactam drugs, such as aminoglycosides, chloramphenicol, quinolones, sulfonamides, tetracyclines and trimethoprim [21, 23].
Chromosomal origin AmpC can occur in two forms: inducible and constitutive. Thus, the exposure of bacteria to β-lactam antibiotics leads to an increase in the production of enzymes. This stimulus may occur at different levels of β-lactamase production, depending on the type of antimicrobial. In both cases, hyperproduction or hyperinduction of the enzyme can be attributed to a mutation in the AmpC regulatory genes [21, 23, 25].
Another important factor to be highlighted is the ability to transfer genes that determine antimicrobial resistance. The genes encoding ESBLs are usually located on plasmids that are highly mobile and can harbor resistance genes to several other unrelated classes of antimicrobials [26]. Carattoli [27] and Zhang [28] identified in Klebsiella pneumoniae isolates, transconjugants of blaCTX-M-type antibiotic resistance genes, blaSHV, blaVIM, blaDHA. Of these, about 67.7% of the transconjugants harbored at least two resistance classes and 32.2% identified three classes of resistance determinants.
The family of SVH-type enzymes appears to be closely related to Klebsiella spp. SHV-1 isthe parent of the SHV-type enzyme class, and most of these enzymes are found in strains of K. pneumoniae [8, 29]. There are a few derivatives of SHV-1, and the changes that occurred in the gene to give rise to its variations were characterized by the substitution of serine for glycine [8].
TEM-1 enzyme is the β-lactamase most commonly found in gram-negative bacteria. Its first derivative is TEM-2, where the only difference is the amino acid substitution from the original β-lactamase. The amino acid substitutions occurring within the TEM enzyme occur in a limited number of positions. Combinations of these amino acid changes result in several subtle alterations in ESBL phenotypes, such as the ability to hydrolyze specific oxyimino-cephalosporins, such as ceftazidime and cefotaxime, or a change in isoelectric points [30]. More than 150 different variants of TEM and SHV that generate resistant have been described in the literature. Among these genes, TEM-3 causes a high rate of resistance concomitant with such antibiotics, being the most reported. The CTX-M enzymes have been recognized as the most prevalent among Enterobacteriaceae and other ESBLs, such as GES, PER, and EBV, have been described as having a lower prevalence [31].
During the 1990s, TEM and SHV were dominant; however, the last decade was marked by the rapid and massive spread of β-lactamases of the CTX-M type, which are currently the most prevalent ESBL in Enterobacteriaceae in Europe and other parts of the world [32]. The existence of the CTX-M enzyme was discovered at the end of the '80s and, more than 100 variations have been identified [33]. Its origin is different from the TEM, and SHV enzymes, the ESBL of CTX-M were acquired by horizontal transfer of genes from other bacteria by translocation or conjugative plasmid, whereas TEM and SHV ESBLs originate from the substitution of the amino acids of parental enzymes [34, 35]. They are classified into five groups based on their amino acid sequence: CTXM-1, CTX-M-2, CTX-M-8, CTX-M-9, and CTX-M-25 [33]. The diversity of CTX-M enzymes from clinical isolates, described today, grossly underestimate the diversity that exists in nature. The rapid global spread of the CTX-M enzyme poses a threat to current antimicrobial therapy strategies, which rely heavily on third-generation cephalosporins to control gram-negative bacterial infections [33].
3.2. Epidemiology
Our results indicate that the population of K. pneumoniae is highly diversified and that in some cases resistant strains seem to have spread throughout the country over time. In addition, we found evidence that some strains independently acquired antimicrobial resistance genes, presumably in response to antimicrobial treatment [5].
The spectrum of infections and the epidemiology of K. pneumoniae changed drastically from the 1970s when this microorganism was identified in a hospital environment. In addition to the considerable colonization efficiency, the antimicrobial resistance allowed K. pneumoniae to spread rapidly in health care settings [14]. The frequency of the association of the bacteria with cases of hospital infections, as well as the high mortality rates caused by K. pneumoniae justify their monitoring [36]. There are reports in the literature of the increasing number of genes associated with the production of ESBLs in K. pneumoniae. Among these, the precursors of SHV and TEM enzymes, as well as CTX-M are isolated in numerous countries, alerting to the occurrence of more than one resistance gene in the same strain [37]. In this study, blaCTX-M-15 and blaSHV-5 genes were identified in 4 of the 7 (57%) of the European countries that were part of this research, both of which were identified in France, Italy, Spain, and Portugal. With featured on France and Spain, where the gene blaCTX-M-15 was isolated twice in rectal swab in both countries (Table 1). Concomitant with the use of antibiotic, the prevalence of Klebsiella pneumoniae strains producing CTX-M-type ESBLs has increased worldwide [46]. The blaCTX-M-15 gene is predominantly present on the European continent due to the spread of isolates initially identified in France and Italy, which have spread to other countries [46].
The β-lactamase CTX-M-15, first detected in 1999 in India, is currently recognized as the most widely distributed CTX-M β-lactamase in the world [46]. CTX-M enzymes, especially CTX-M-15, are related to various epidemiological situations and have spread throughout the continents due to epidemic plasmids and/or epidemic strains [47]. Corroborating this statement, in the present study, the blaCTX-M-15 gene was found in K. pneumoniae in all continents evaluated Table(1-4). Many of the emerging antimicrobial resistance problems of the nineties have been characterized by difficulty in recognition of resistance in the laboratory, particularly by rapid susceptibility-test methods. The emergence of plasmid-encoded ESBLs is a significant evolution in antimicrobial resistance. Outbreaks due to the dissemination of ESBL-producing bacterial strains and to the dissemination of ESBL-encoding plasmids among different species of the family Enterobacteriaceae have been described in hospitals and other health care facilities [46, 47].
Some authors identified antimicrobial resistance genes in K. pneumoniae and demonstrated the CTX-M-15 and SHV-5 domain isolated from Portuguese hospitals [45]. The production of the SHV-type ESBLs in K. pneumoniae is associated with an increased tendency to invade epithelial cells and expression of fimbrial adhesions [48]. The propagation of these genes is influenced by the amplification of the epidemic and international clones of K. pneumoniae and specific types of plasmids. The same authors also highlight the potential for antibiotic resistance and virulence plasmid dissemination [45]. The origin of the blaCTX-M-15 gene identified in K. pneumoniae in Portugal is clone ST336, and the most recent occurrence in France, described in the literature, reported a clonal outbreak involving K. pneumoniae isolates recovered from a single hospital in the Picardie region of northern France [45, 49]. This outbreak was caused by a resistance to colistin with OXA-48 and CTXM-15 which produced K. pneumoniae type ST11 that was susceptible only to cefoxitin. The first identification of the ST11 clone in K. pneumoniae occurred in 1997 in France [49]. This clone has been associated to the acquisition of resistance genes to a broad spectrum of antimicrobial agents, as well as to the dissemination of OXA-48, in addition to VIM (Verona Imipenemase), NDM (New Delhi Metallo-β-lactamase) and KPC [40, 49].
The most recent occurrence of blaCTX-M-15 in other European countries was described by Rodrigues and Brañas [40, 45]. The first author related finding of the increased incidence of CTX-M-15, SHV and several other types ESBL were observed in a Portuguese hospital associated with the increase of MDR K. pneumoniae epidemic clones, the second reported blaOXA-48 and blaCTX-M-15 as responsible for a major outbreak involving 44 patients at a hospital in Madrid, Spain, 2009 to 2014 [40, 45].
In the United Kingdom and Sweden, the only resistance genes identified were NDM-type, with blaNDM-1 being isolated in both countries (Table 1). Among the NDM isolates identified in the United Kingdom, ST231 was the most common type along with ST147 and ST273. In the case of Sweden, the isolates belonged to four different sequence types, as they were imported from various regions of the world such as India and Iraq [43]. Strains of K. pneumoniae NDM-1 positive can exhibit relatively high clonal diversity. In Sweden and the United Kingdom clone ST14 was the most frequently observed type. Clone K. pneumoniae ST14 has been described as a host for the NDM-1 enzyme and is also a frequent host of CTX-M enzymes. In addition, ST14 is a single locus variant (SLV) of ST15, which often encodes the CTX-M type ESBLs. The production of the NDM-1 enzyme by isolates belonging to clone ST11 and complex ST147-ST273 is very relevant information since clone ST11 is a frequent host of CTX-M and KPC and is also an SLV of ST258, making it fundamental for its dissemination [43, 50].
Of the European countries correlated in this study, Spain stands out as the site where the largest number of different genes (15 in total) was identified, which shows the genetic diversity in relation to the occurrence of K. pneumoniae with resistance to the antibiotic β-lactams (Table 1). This characteristic, from the point of view of the therapeutic of the infection, is a difficult factor, considering that the variation of resistance genes decreases the antimicrobial therapy options.
The country where the lowest gene diversity occurred was Greece, where the only isolated gene was blaKPC-2 in two outbreak cases (Table 1). Considering that the outbreaks were of monoclonal origin and that there was no identification of a common source or an environmental reservoir, the main dissemination mechanism of the blaKPC-2 in these cases was the transmission from patient to patient [41, 42]. The smallest genetic diversity in Greece, identified in this study, may be related to a low number of publications in this country.
| Europe | |||
|---|---|---|---|
| Country | Genes (N. reports)* |
Isolated (N. reports)* |
References |
| France | blaCTX-M-15 (2) | Rectal swab (2) Catheter Urine Wound exudate Respiratory tract |
(Baraniak et al. 2013)38 |
| blaCTX-M-1 | |||
| blaCTX-M-40 | |||
| blaSHV-5 | |||
| blaSHV-12 | |||
| blaSHV-2 | |||
| blaTEM-3 | |||
| blaCMY-2 | |||
| blaOXA-48 | |||
| Italy | blaCTX-M-2 | Rectal swab | (Baraniak et al. 2013)38 |
| blaCTX-M-15 | |||
| blaTEM-3 | |||
| blaCTX-M-1 | |||
| blaSHV-5 | |||
| Spain | blaCTX-M-1 | Bloodstream Bronchial aspirates Clinical isolates Non-surgical wound Rectal swabs (2) Sputum Surgical wound Urine Vascular catheters |
(Hernández et al. 2005)39 (Baraniak et al. 2013)38 (Brañas et al. 2015)40 |
| blaCTX-M-3 | |||
| blaCTX-M-9 | |||
| blaCTX-M-10 | |||
| blaCTX-M-15(2) | |||
| blaSHV-2 (2) | |||
| blaSHV-2a | |||
| blaSHV-5 | |||
| blaSHV-12 | |||
| blaDHA-1 | |||
| blaTEM-3 | |||
| blaTEM-4 | |||
| blaTEM-25 | |||
| blaTEM-133 | |||
| blaOXA-48 | |||
| blaVIM-1 | |||
| Greece | blaKPC-2 (2) | Respiratory tract Catheter tip surgical site Bloodstream (2) Feces Bronchial secretions Pus Central venous catheter |
(Maltezou et al. 2009)41 (Souli et al. 2010)42 |
| United Kingdom |
blaNDM blaNDM-1 |
Urine (2) Respiratory tract |
(Giske et al. 2012)43 (Jain et al. 2014)44 |
| Portugal |
blaCTX-M-15 blaSHV-12 blaSHV-106 blaSHV-28 blaSHV-55 blaSHV-2 blaSHV-5 |
Urine Bloodstream Exudate Sputum |
(Rodrigues et al. 2014)45 |
| Sweden |
blaKPC blaNDM-1 |
Urine Respiratory tract |
(Giske et al. 2012)43 |
In Asia and Oceania, 41 different resistance gene types have been identified in the literature. Of these, the greatest diversity occurred in China, where 14 of the 41 genes were described, corresponding to 41% of the total of related genes in Table 2. The variety of genes in China can be justified by its high population density, associated with intense migrations.
The second country with the greatest diversity of genes was Israel (12 different genes), followed by Turkey (11 genes) and Iran (10 genes). The lowest variety of β-lactam resistance genes occurred in Pakistan (2 genes), according to data in Table 2. Despite the lower diversity, the genes identified in Pakistan, blaKPC, and blaNDM-1, are of great importance for tracking the control of microbial resistance. The first gene is characterized by its wide plasmodial spread, and the second has broad relationships with the globalization and population mobility [63, 64]. The blaNDM-1 gene was first identified in 2009 in a Swedish patient of Indian origin, who acquired a urinary tract infection in New Delhi, caused by a strain of K. pneumoniae that showed resistance to β-lactam antibiotics [64, 65].
| Asia and Oceania | |||
|---|---|---|---|
| Country | Genes | Isolated | References |
| (N. reports)* | (N. reports)* | ||
| China | blaCMY-2 | Bloodstream | (Giske et al. 2012) [43] |
| blaCTX-M | Urine (2) | (Ginn et al. 2014) [51] | |
| blaCTX-M-1 | Respiratory tract | (Qi et al. 2015) [52] | |
| blaCTX-M-15 | |||
| blaCTX-M-9 | |||
| blaDHA | |||
| blaIMP-4 | |||
| blaKPC | |||
| blaKPC-2 | |||
| blaNDM-1 | |||
| blaOXA-10 | |||
| blaSHV-12 | |||
| blaSHV-5 | |||
| blaTEM | |||
| Israel | blaCMY-2 | Swab retal | (Leavitt et al. 2007) [53] |
| blaCTX-M-10 | Urine | (Baraniak et al. 2013) [58] | |
| blaCTX-M-14 | Body fluids | ||
| blaCTX-M-2 | Wounds | ||
| blaCTX-M-39 | Catheter tips | ||
| blaKPC-2 | Bloodstream | ||
| blaKPC-3 | Respiratory tracts | ||
| blaOXA-4 | |||
| blaSHV-12 | |||
| blaSHV-27 | |||
| blaSHV-5 | |||
| blaTEM-1 | |||
| blaTEM-3 | |||
| India | blaNDM-1 | SENTRY | (Castanheira et al. 2011) [54] |
| blaOXA-181 | Antimicrobial | ||
| blaVIM-5 | Surveillance Program | ||
| (hospital patients) | |||
| Singapore | blaCMY-2 | Bloodstream | (Ginn et al. 2014) [51] |
| blaCTX-M-1 | Urine | ||
| blaCTX-M-9 | |||
| blaDHA | |||
| blaSHV-12 | |||
| blaSHV-5 | |||
| blaTEM | |||
| Australia | blaCTX-M-3 | Bloodstream | (Paterson et al. 2003) [55] |
| blaSHV-2 | |||
| blaTEM-1B | |||
| Turkey | blaCTX-M-2(2) | Bloodstream (4) | (Paterson et al. 2003) [55] |
| blaCTX-M-15 | Urine (3) | (Bali et al. 2010) [56] | |
| blaPER-1 | Sputum | (Carrer et al. 2010) [57] | |
| blaSHV-2 (2) | Wound (3) | ||
| blaSHV-5 (2) | Abscesses | ||
| blaSHV-12 (3) | Catheter (3) | ||
| blaTEM-1B | Peritoneum fluid | ||
| blaTEM-1 (2) | Cerebrospinal fluid | ||
| blaTEM-2 | Endotracheal aspirate (2) | ||
| blaOXA-48 | Pus (2) | ||
| blaOXA-9 | Ascites (2) | ||
| blaOXA-1 | |||
| Taiwan | blaCTX-M-14 | Nationwide | (Ling Ma et al. 2009) [58] |
| blaCTX-M-15 | surveillance | ||
| blaCTX-M-3 | program | ||
| blaSHV-1 | |||
| blaSHV-11 | |||
| blaSHV-12 | |||
| blaTEM-31 | |||
| Iran | blaCTX-M | Abscess | (Nasehi et al. 2010) [59] |
| blaCTX-M-15 | Abdominal exudates | (Feizabadi et al. 2010) [60] | |
| blaPER | Bloodstream (2) | ||
| blaSHV | Bronchial | ||
| blaSHV-1 | Eyes exudates | ||
| blaSHV-12 | Sputum | ||
| blaSHV-31 | Stool | ||
| blaTEM | Trachea | ||
| blaTEM-1 | Umbilical exudates | ||
| blaTEM-79 | Urine (2) | ||
| Wound (2) | |||
| Lebanon | blaCTX-M-15 | Stool sample | (Tokajian et al. 2015 [62]) |
| blaOXA-1 | |||
| blaSHV-11 | |||
| blaTEM-1b | |||
| Pakistan | blaKPC-2 | Pus | (Sattar et al. 2016) [61] |
| blaNDM-1 | Urine | ||
| Catheter tip | |||
| Sputum | |||
| Wound | |||
Of the isolates of K. pneumoniae from Singapore, the occurrence of blaCTX type genes predominated, with the blaCTX-M-1 group being more numerous than the others. Literature data report transport of methylase genes in isolated MDR K. pneumoniae strains in Chinese hospitals [52]. In most countries of Asia and Oceania, the blaCTX-M gene, in particular, blaCTX-M-15, was detected in K. pneumoniae, as shown in Table 2. A study published in 2015 reported, for the first time, the isolation of K. pneumoniae CTX-M-15, SHV-11, and KPC belonging to ST336 in Lebanon [62]. The contributors to the dissemination of the blaCTX-M-15 gene highlight the clonal complex 17, among them ST336 [45] is related to the presence of this gene in clinical isolates in several countries of the world. In the present study, the occurrence of the blaCTX-M-15 gene is reported in all continents evaluated, except in the African continent, revealing its endemicity and, highlighting the dissemination of this gene at a global level through human mobility. This clone was isolated for the first time from a patient (nurse), 62 years old, with an abdominal tumor and who had recurrent urinary infections. The ST336 epidemic clone belongs to CG17, and studies revealed the prevalence of this strain in France, Germany, Italy, USA and Greece [38, 66]
In this study, K. pneumoniae with the presence of the blaKPC gene was isolated in all the evaluated continents, as described in Tables 1, 2, 3 and 4. Although it occurs in all continents evaluated, the blaKPC-2 type gene was more prominent in the Asian and American continents (Tables 2 and 3), and is described as the second most frequent gene. The most prevalent β-lactamase in the family Enterobacteriaceae is the type of KPC, which was found in Klebsiella pneumoniae, especially in the United States, Asia, United Kingdom, Israel and Europe [65]. The propagation of the blaKPC gene to other countries is associated with cases imported from countries where they are endemic [66]. Besides, population mobility is recognized as a major factor in the spread of microorganisms resistant to antimicrobial drugs. Thus, the emergence of K. pneumoniae producing blaKPC in the United States in 2001 may be associated with the onset of travel-related outbreaks in other countries [63].
In the literature, the first identification of the blaOXA-48 gene in a K. pneumoniae isolate occurred in Istanbul, Turkey, in 2001, since then the β-lactamase OXA-48 has become an important carbapenemase associated with the Entero bacteriaceae family in Europe, North Africa, and Asia [35, 72].
In the Americas, the highest diversity of β-lactam resistance genes occurred in Brazil, where 12 of the 25 different types of genes identified on the continent were reported, representing 48% of the total genes present in this continent (Table 3). Secondly, in the diversity of resistance genes, the United States of America (USA) presented 10 of the 25 genes identified; followed by Canada (9 different genes) and Argentina (6 different genes). The lowest resistance gene diversity occurred in Mexico (3 different genes), according to data described in Table 3. The smallest variety of genes in Mexico can be justified by the low number of publications on the subject in this country.
The occurrence of β-lactamase gene diversity produced by K. pneumoniae may be closely related to the variety of antimicrobials used in the treatment of infection over the decades. With the evolution of antimicrobial resistance in bacteria, it is necessary to continually re-evaluate the laboratory detection of ESBLs and potential treatment options for organisms that produce ESBLs [1].
Among the resistance genes identified in the American continent, blaCTX-M-2, blaSHV-1, blaSHV-5, and blaSHV-2 were isolated from K. pneumoniae in 60% of the American countries listed in this study. The genes blaCTX-M-15, blaCTX-M-1, blaSHV-11, blaSHV-12, blaTEM-1, blaTEM-10 and blaKPC-2 were the second most frequent in the Americas, being found in 40% (Table 3).
TEM-10 was ESBL type TEM which is most frequently detected in K. pneumoniae isolates, being found in different geographic regions (United States, South Africa, and Argentina). The identification of genes encoding the production of TEM-10 β-lactamase isolated from K. pneumoniae in Argentina in 2003 represented the first report of ESBL-type TEM-10 in South America [55]. In Argentina, most ESBLs belong to the CTX-M class, referring to a growing family of ESBLs that shares a single phenotype that confers in vitro resistance to cefotaxime, ceftriaxone and many times to cefepime [74].
| America | |||
| Country | Genes | Isolated | References |
| (N. reports)* | (N. reports)* | ||
| Argentina | blaTEM-10 | Bloodstream (2) | (Paterson et al. 2003) [55] |
| blaSHV-5 | Surgical site | (Pasteran et al. 2008) [67] | |
| blaSHV-2 | |||
| blaTEM-12 | |||
| blaCTX-M-2 | |||
| blaKPC-2 | |||
| Brazil | blaKPC-2 (2) | Bloodstream (3) | (Peirano et al.2009) [68] |
| blaSHV-1 | Urine (3) | (Pereira et al. 2013) [69] | |
| blaSHV-11 | Abdominal fluid | (Tollentino et al. 2011) [70] | |
| blaSHV-12 (2) | Respiratory tract secretion | (Andrade et al. 2014) [48] | |
| blaSHV-31 | Tracheal secretion (2) | ||
| blaSHV-38 | Venous catheter | ||
| blaCTX-M-2 (2) | |||
| blaCTX-M-15 | |||
| blaOXA-9 | |||
| blaTEM-1(2) | |||
| blaCTX-M-1 | |||
| blaCTX-M-8 | |||
| Canada | blaCTX-M-2 | Bloodstream | (Denisuik et al. 2013) [71] |
| blaSHV-11 | Urine | ||
| blaCTX-M-3 | Respiratory tract | ||
| blaCTX-M-14 | Secretions | ||
| blaSHV-1 | |||
| blaTEM-1 | |||
| blaCTX-M-15 | |||
| blaOXA-1 | |||
| blaSHV-2 | |||
| Mexico | blaSHV-1 | Bloodstream | (Miranda et al. 2004) [73] |
| blaSHV-2 | Cerebrospinal fluid | ||
| blaSHV-5 | |||
| United States | blaCTX-M-1 | Bloodstream | (Ginn et al. 2014) [51] |
| blaCTX-M-9 | Urine | (Mathers et al. 2013) [72] | |
| blaSHV | Perirectal swab (2) | ||
| blaSHV-5 | Fluid from abdominal drain | ||
| blaSHV-12 | |||
| blaTEM-10 | |||
| blaTEM (2) | |||
| blaOXA-10 | |||
| blaOXA-48 | |||
| blaOXA-181 | |||
In Brazil, the first reported case of KPC occurred in 2009. Data from a survey published in 2005 pointed out that the number of ESBL positive strains is increasing worldwide from 30% to 60% of strains of Klebsiella [37]. A major concern emerged in Brazil after the isolation and identification of Klebsiella pneumoniae with concomitant expression of MBL IMP-1 and ESBL CTX-M, since the production of these enzymes together gives the microorganism resistance to all antibiotics available for treatment, including carbapenems [78]. A study on the antimicrobial resistance of Klebsiella spp., in Brazil, reported the trend of resistance growth in most of the antibiotics tested, indicating the need for new studies to monitor both the evolution of resistance and the emergence of new resistance mechanisms. The rapid spread of ESBL-producing K. pneumoniae is a major clinical and public health concern. These broad-spectrum β-lactamases are increasing in new locations around the world, indicating this to be a continuous process [79].
The literature data are still scarce in reporting the problem of ESBLs produced by K. pneumoniae in Africa, especially in sub-Saharan Africa [65, 80]. In the present study, the presence of K. pneumoniae presenting blaNDM-1 resistance gene, as described in (Table 4), was identified in Kenya and South Africa. NDM-1 is a type of MBL whose denomination is related to the city of origin, being a common practice in the case of transferable MBLs [81]. Isolates from the Enterobacteriaceae family harboring the blaNDM-1 gene are responsible for causing a variety of infections including urinary tract infections, septicemia, pulmonary infections, diarrhea, peritonitis, device-associated infections, and soft tissue infections [82]. The blaNDM-1 gene is a genetic element found in different plasmids. This gene is characterized by a large capacity of duplication migrating from bacterium to bacterium, which explains its rapid dissemination [64]. Also, blaNDM-1 is easily disseminated between different bacterial species by lateral transfer. These characteristics make globalization a favorable aspect of gene dissemination, where travel represents a high risk of a global pandemic among Enterobacteriaceae. Since these plasmids also harbor genes that confer resistance to almost all antibiotics, making rapid their spread in clinically relevant bacteria a serious threat to therapy [64, 81]. The results of the present study report the occurrence of the K. pneumoniae isolated blaNDM-1 gene in all continents, except in the Americas (Table 3).
Although the blaNDM-1 gene isolated from K. pneumoniae in the American continent was not related to the present study (because the articles found did not meet the inclusion criteria), a study identified the first isolate of this gene in South America [83]. However, data from the literature mainly demonstrate the presence of the gene in the Americas isolated from other enterobacteria other than K. pneumoniae. This occurrence demonstrates the ease of dissemination of this gene among different bacterial species by lateral transfer, especially in the Enterobacteriaceae family, as previously described [84].
The propagation of NDM-1 shows some problems related to the occurrence of MBLs, which include: the absence of a routine standardized phenotypic test for MBL detection; the consequent high prevalence of unrecognized asymptomatic carriers, allowing international dissemination of these bacteria; the scarcity of effective antibiotics available today; and the possibility of their spread in many different gram-negative bacteria [81]. In addition to NDM-1 β-lactamase, other resistance genes have been identified in South Africa, namely: blaSHV-5, blaSHV-2, blaTEM-10, blaTEM12, blaCTX-M-2 and blaKPC-2, as described in (Table 4). In 2012, the occurrence of K. pneumoniae in South Africa with reduced susceptibility to carbapenems due to ESBL CTX-M was reported. In this study, the blaCTX-M-2 gene was identified in South Africa [55, 76]. However, there was no isolation in Kenya (Table 4), as well as other types of CTX-M, originating from K. pneumoniae were not reported.
4. CONCLUSION
The content of the present study characterizes the relevance/importance of epidemiological knowledge related to mechanisms of resistance of β-lactam antibiotics associated with the microorganism Klebsiella pneumoniae from the analysis of scientific production. One of the results of great importance in this study is the coexistence of different classes of β-lactamases in the same strain, representing not only a therapeutic challenge but also a challenge in the diagnosis of infectious disease. Among the microbial resistance genes originating from K. pneumoniae and identified in this study, blaSHV-12 – which codes one of the most prevalent ESBLs - was the most frequent in Americas, Europe, and Asia, except in Africa. Otherwise, blaCTX-M-2 and blaSHV-5 genes, the second most reported, were found in all continents, characterizing their cosmopolitan distribution. The highest genetic diversity was observed in Asian and Oceania continents, where 41 different types of genes were reported. Additionally, the verification of resistance genes shows a variation over time and location, which highlights the importance of evaluating the mechanisms or strategies by which these variations occur. Finally, understanding the occurrence and distribution of these genes may generate possibilities of future interference in this process with the production of new drugs which are more efficient and specific, thereby providing results that allow assisting the health team in the most appropriate therapy.
| Africa | |||
| Country | Genes (N. reports)* |
Isolated (N. reports)* |
References |
| Kenya | blaNDM-1 | Urine Urethral pus |
(Poirel et al. 2011) [75] |
| South Africa |
blaTEM-10 blaSHV-5 blaSHV-2 blaTEM-12 blaCTX-M-2 blaNDM-1 blaKPC-2 |
Bloodstream Urine Sputum |
(Paterson et al. 2003) [55] (Brink et al. 2012) [76] |
| Tanzania |
blaCTX-M-15 blaTEM-104 blaSHV-11 blaTEM-176 |
Bloodstream Wound swab Urine Pus |
(Mshana et al. 2013) [77] |
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
This work was supported by Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro process no. E-26/201.577/2018 and E-26/201.185/2014. Conselho Nacional de Desenvolvimento Científico e Tecnológico (process no. 311422/2016-0, CNPq, Brazil).
CONFLICT OF INTEREST
The author declares no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.